Overview
MLGenX LLM Perturbation Challenge
The next wave of AI for science may emerge from the intersection of two frontier ideas: LLMs/agents as general reasoning systems, and virtual cells as predictive models of biological response. The MLGenX LLM Perturbation Challenge is designed to explore exactly that boundary.
In this challenge, participants tackle the focused but scientifically meaningful task of predicting the outcome of a Perturb-seq experiment: given a knockdown of gene X, predict whether a target gene Y will be up-regulated, down-regulated, or unchanged in activated macrophages. This framing turns perturbation biology into a benchmark for machine reasoning: can an AI system combine biological knowledge, causal intuition, and structured inference to anticipate the downstream effects of intervention?
The challenge is a controlled comparison of three emerging paradigms for scientific LLMs and agentic reasoning: prompt-guided reasoning, agentic tool use, and task-specific fine-tuning. In that sense, BioReasoningChallenge is not just a competition in predictive performance. It is a benchmark for a broader question shaping the future of computational biology: can frontier LLMs and agents become practical building blocks for virtual cell models?

Tasks – Perturbation Prediction
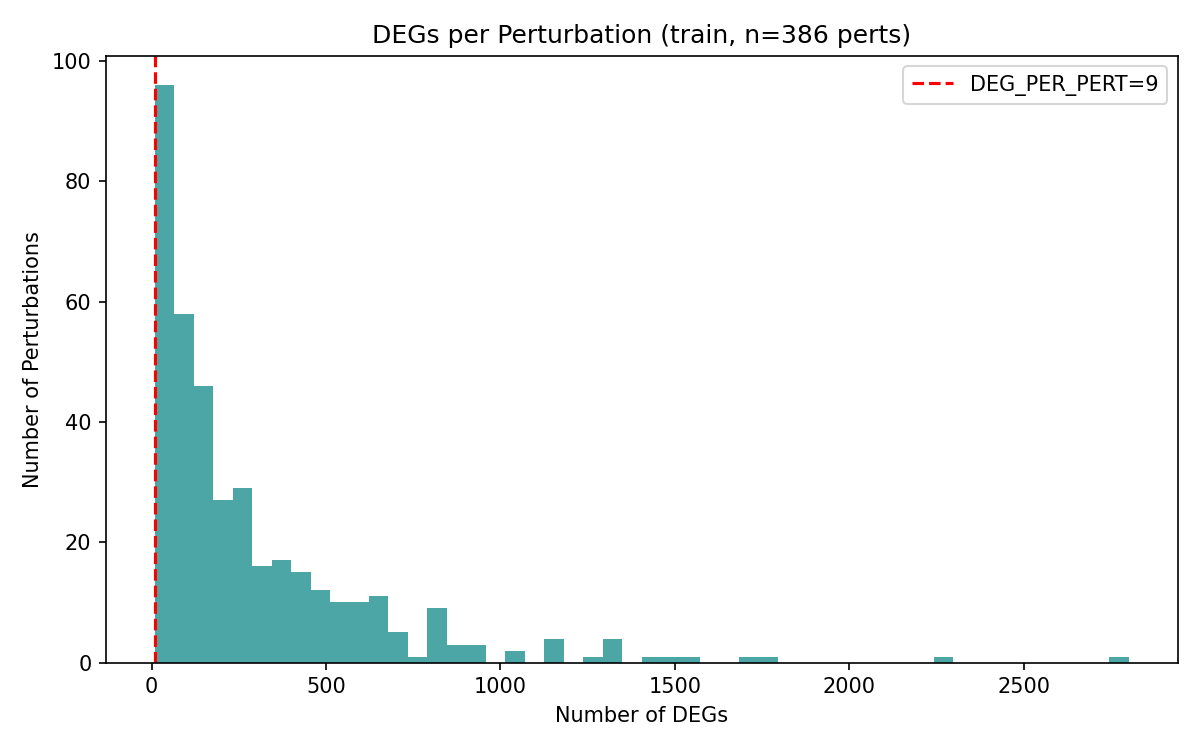
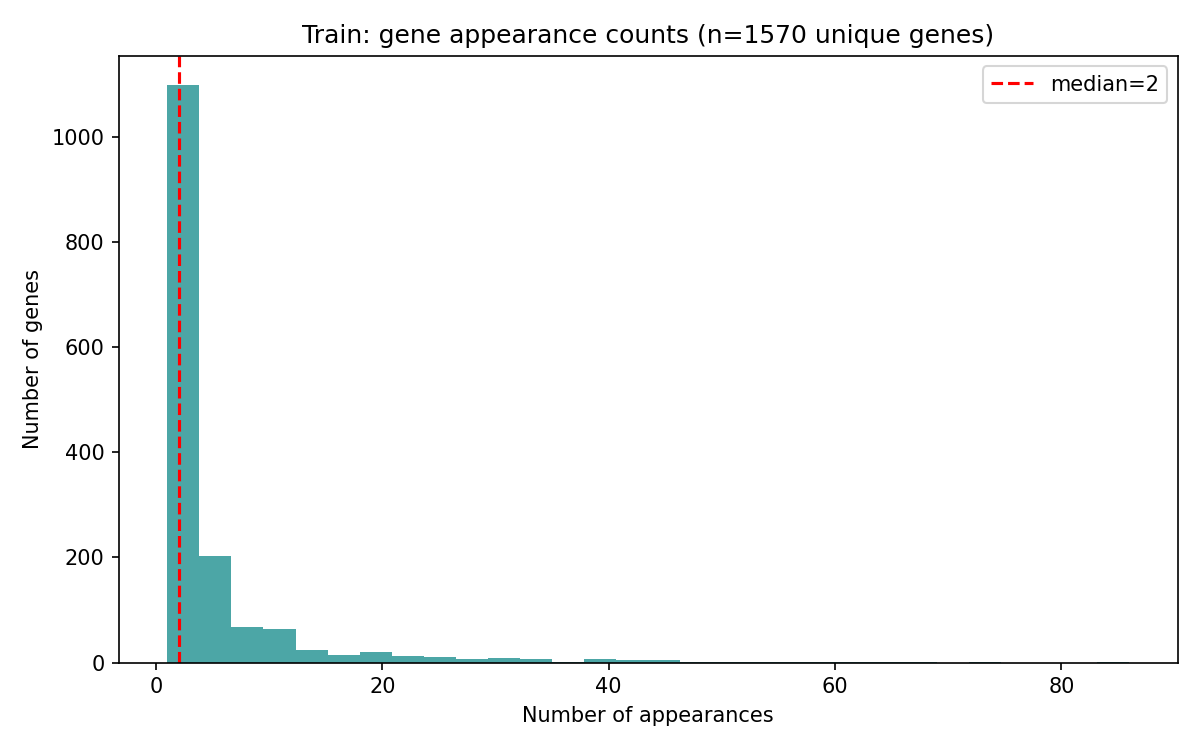
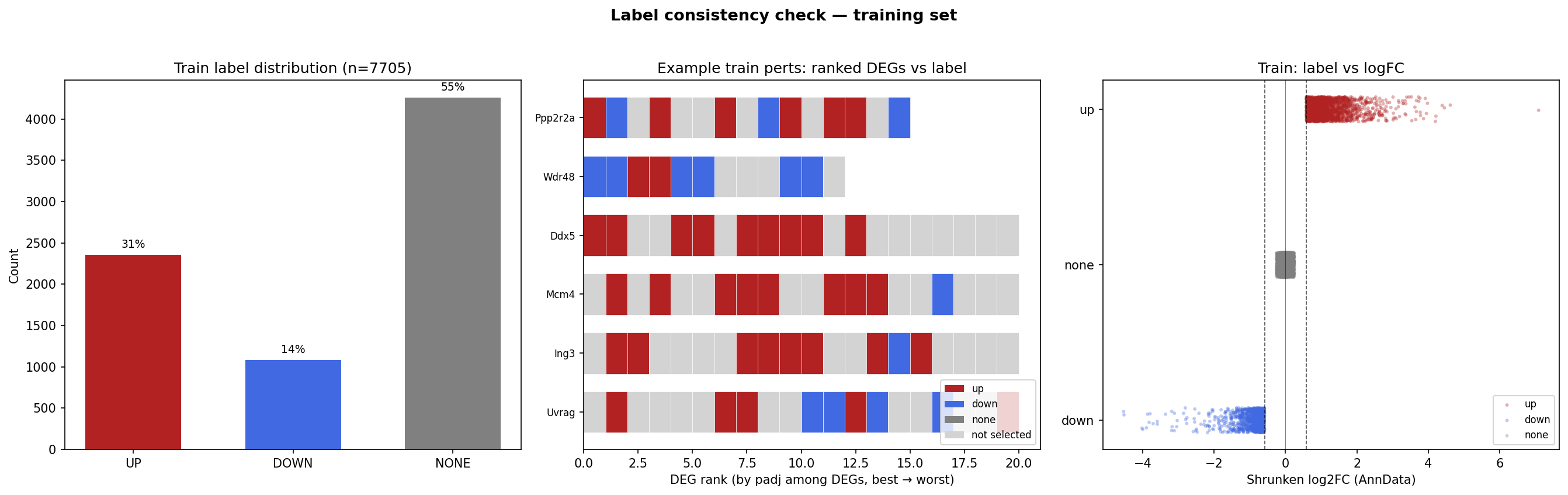
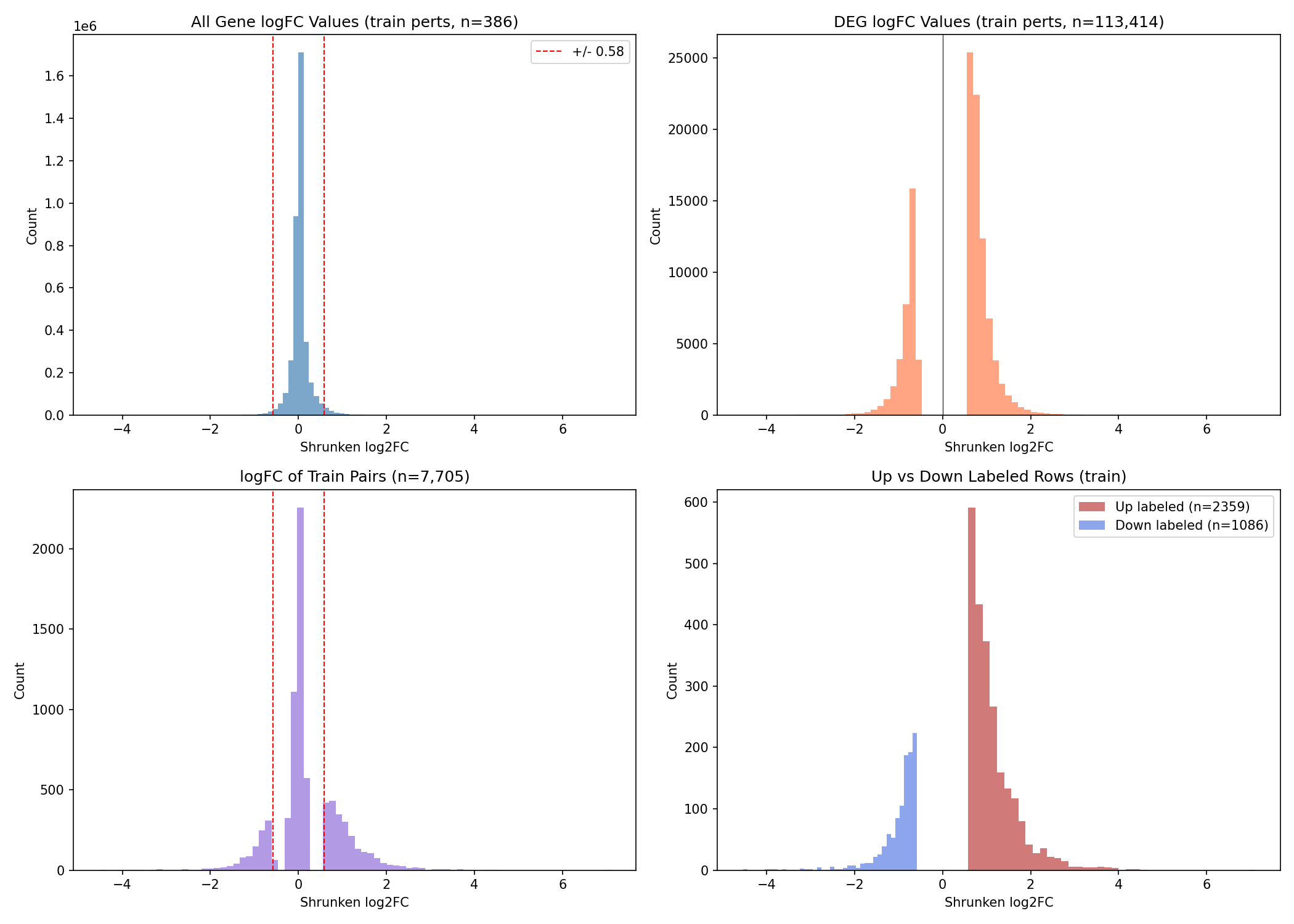
The core task is a supervised predictive challenge where participants must infer the effect of a specific gene perturbation on the expression of a target gene. Structurally, the task is defined as: Perturbation gene X → gene Y up/down/no-change. Participants will be scored based on their prediction of these post-processed labels, which indicate whether gene Y is up-regulated, down-regulated, or unchanged following the perturbation of gene X.