(Advanced) Using gReLU with external pytorch models#
This tutorial shows how to take a pytorch model that was not trained using gReLU, and make it compatible with gReLU’s downstream functions (e.g. inference, variant effect prediction, interpretation and design).
import numpy as np
import pandas as pd
import torch
from torch import nn
import os
Load an external pytorch model#
We will load the pytorch version of the Basset model from the Kipoi repository (https://kipoi.org/models/Basset/). You can also try this with any other pytorch sequence-to-function model.
Kipoi provides us the following important information about the model:
This is the Basset model published by David Kelley converted to pytorch by Roman Kreuzhuber. It categorically predicts probabilities of accesible genomic regions in 164 cell types (ENCODE project and Roadmap Epigenomics Consortium). Data was generated using DNAse-seq. The sequence length the model uses as input is 600bp. The input of the tensor has to be (N, 4, 600, 1) for N samples, 600bp window size and 4 nucleotides. Per sample, 164 probabilities of accessible chromatin will be predicted.
You will need to uncomment the cell below to install kipoi.
#!pip install kipoi
import kipoi
kipoi_model = kipoi.get_model('Basset', with_dataloader=False)
kipoi_model = kipoi_model.model.cpu()
Downloading https://zenodo.org/record/1466068/files/pretrained_model_reloaded_th.pth?download=1 to /home/lala8/.kipoi/models/Basset/downloaded/model_files/weights/4878981d84499eb575abd0f3b45570d3
We can view the structure of this model below:
kipoi_model
Sequential(
(0): Conv2d(4, 300, kernel_size=(19, 1), stride=(1, 1))
(1): BatchNorm2d(300, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU()
(3): MaxPool2d(kernel_size=(3, 1), stride=(3, 1), padding=0, dilation=1, ceil_mode=False)
(4): Conv2d(300, 200, kernel_size=(11, 1), stride=(1, 1))
(5): BatchNorm2d(200, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(6): ReLU()
(7): MaxPool2d(kernel_size=(4, 1), stride=(4, 1), padding=0, dilation=1, ceil_mode=False)
(8): Conv2d(200, 200, kernel_size=(7, 1), stride=(1, 1))
(9): BatchNorm2d(200, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(10): ReLU()
(11): MaxPool2d(kernel_size=(4, 1), stride=(4, 1), padding=0, dilation=1, ceil_mode=False)
(12): Lambda()
(13): Sequential(
(0): Lambda()
(1): Linear(in_features=2000, out_features=1000, bias=True)
)
(14): BatchNorm1d(1000, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(15): ReLU()
(16): Dropout(p=0.3, inplace=False)
(17): Sequential(
(0): Lambda()
(1): Linear(in_features=1000, out_features=1000, bias=True)
)
(18): BatchNorm1d(1000, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(19): ReLU()
(20): Dropout(p=0.3, inplace=False)
(21): Sequential(
(0): Lambda()
(1): Linear(in_features=1000, out_features=164, bias=True)
)
(22): Sigmoid()
)
Modify the model to make it gReLU-compatible#
Step 1: Check the input and output formats#
First, we need to pay attention to the shapes of the model’s input and output shapes. As per Kipoi:
The input of the tensor has to be (N, 4, 600, 1) for N samples, 600bp window size and 4 nucleotides.
Kipoi doesn’t tell us the shape of the output tensor produced by this model, so let’s check by creating a random tensor of the required shape and passing it through the model.
random_tensor = torch.rand(2, 4, 600, 1) # N=2
random_tensor.shape
torch.Size([2, 4, 600, 1])
output_shape = kipoi_model(random_tensor).shape
print(output_shape)
torch.Size([2, 164])
Step 2: Define adapter layer(s) if necessary#
We see that this model expects inputs of shape (N, 4, input_len, 1) and produces outputs of shape (N, T). Here N is the number of examples, input_len is the sequence length and T is the number of tasks.
However, gReLU models take inputs of shape (N, 4, input_len) and produce outputs of shape (N, T, output_len). For binary classification models like Basset, the output_len is 1, so outputs should be of shape (N, T, 1).
Hence, we will need to add adapter layers to this model to create inputs and outputs of the right format. Specifically:
gReLU inputs of shape
(N, 4, input_len)should be converted to the model’s required shape(N, 4, input_len, 1). This requires adding a new axis.The model’s outputs of shape
(N, T)should be converted to gReLU’s required shape(N, T, 1). This also requires adding a new axis.
We can achieve both of these by creating an AddNewAxis layer, as below:
class AddNewAxis(nn.Module):
def __init__(self):
super().__init__()
def forward(self, x):
return x.unsqueeze(-1)
Step 3: Construct embedding, head and activation sections#
We see that kipoi_model consists of a sequence of 21 pytorch layers plus the sigmoid activation. All gReLU models are separated into three parts: the embedding, the head, and the final activation.
Generally, all the layers upto the final linear layer are the embedding. The last linear layer of the model, which produces the right number of outputs (164 in this case) is the head. The final non-linear layer of the model, in this case Sigmoid() is the activation.
We now split kipoi_model into embedding, head, and activation sections:
embedding = kipoi_model[:21]
head = kipoi_model[21]
activation = kipoi_model[22]
We can look at these individually:
embedding
Sequential(
(0): Conv2d(4, 300, kernel_size=(19, 1), stride=(1, 1))
(1): BatchNorm2d(300, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU()
(3): MaxPool2d(kernel_size=(3, 1), stride=(3, 1), padding=0, dilation=1, ceil_mode=False)
(4): Conv2d(300, 200, kernel_size=(11, 1), stride=(1, 1))
(5): BatchNorm2d(200, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(6): ReLU()
(7): MaxPool2d(kernel_size=(4, 1), stride=(4, 1), padding=0, dilation=1, ceil_mode=False)
(8): Conv2d(200, 200, kernel_size=(7, 1), stride=(1, 1))
(9): BatchNorm2d(200, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(10): ReLU()
(11): MaxPool2d(kernel_size=(4, 1), stride=(4, 1), padding=0, dilation=1, ceil_mode=False)
(12): Lambda()
(13): Sequential(
(0): Lambda()
(1): Linear(in_features=2000, out_features=1000, bias=True)
)
(14): BatchNorm1d(1000, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(15): ReLU()
(16): Dropout(p=0.3, inplace=False)
(17): Sequential(
(0): Lambda()
(1): Linear(in_features=1000, out_features=1000, bias=True)
)
(18): BatchNorm1d(1000, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(19): ReLU()
(20): Dropout(p=0.3, inplace=False)
)
head
Sequential(
(0): Lambda()
(1): Linear(in_features=1000, out_features=164, bias=True)
)
activation
Sigmoid()
Step 4: Format the embedding and head#
We need to add the adapter layer we defined previously to the embedding and head, to ensure that they produce the correct input and output shape.
embedding = nn.Sequential(AddNewAxis(), *[l for l in embedding])
head = nn.Sequential(*[l for l in head], AddNewAxis())
The head in a gReLU model also needs to have an attribute n_tasks. This is the number of outputs of the model:
head.n_tasks = 164
Wrap the model in a LightningModel object#
We can construct a LightningModel object using the embedding, head and activation objects.
from grelu.lightning import LightningModel
grelu_model = LightningModel(model_params={'model_type':'BaseModel', 'embedding':embedding, 'head':head})
grelu_model.activation = activation
Add the task metadata#
We can also add the metadata concerning the 164 cell types that the model is trained on.
tasks = pd.read_table('https://raw.github.com/davek44/Basset/refs/heads/master/data/models/targets.txt', header=None,
names=['name', 'source'])
print(len(tasks))
tasks.head()
164
| name | source | |
|---|---|---|
| 0 | 8988T | encode/wgEncodeAwgDnaseDuke8988tUniPk.narrowPe... |
| 1 | AoSMC | encode/wgEncodeAwgDnaseDukeAosmcUniPk.narrowPe... |
| 2 | Chorion | encode/wgEncodeAwgDnaseDukeChorionUniPk.narrow... |
| 3 | CLL | encode/wgEncodeAwgDnaseDukeCllUniPk.narrowPeak.gz |
| 4 | Fibrobl | encode/wgEncodeAwgDnaseDukeFibroblUniPk.narrow... |
grelu_model.data_params['tasks'] = tasks.to_dict(orient="list")
Test the model#
Let’s make sure that the model can generate predictions for a random sequence:
from grelu.sequence.utils import generate_random_sequences
test_input = generate_random_sequences(n=1, output_format='strings', seq_len=600, seed=0)
test_output = grelu_model.predict_on_seqs(test_input)
test_output.shape, test_output.min(), test_output.max()
((1, 164, 1), np.float32(0.13277043), np.float32(0.72227246))
Use the model#
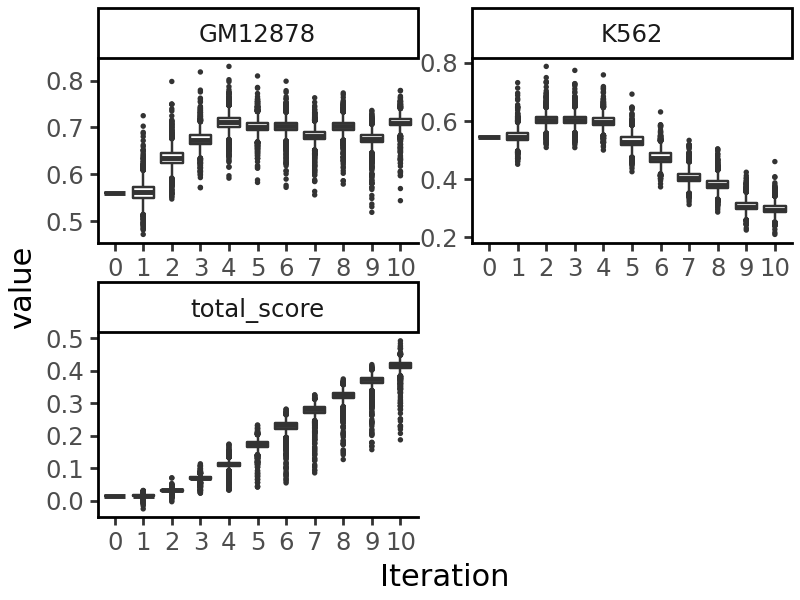
We can now use this model for various downstream functions. As an example, we will take the random sequence we generated above, and edit it using directed evolution to produce a sequence accessible in GM12878 but not in K562 cells. See Tutorial 4 for more details on the directed evolution process.
from grelu.transforms.prediction_transforms import Specificity
gm12878_specificity = Specificity(
on_tasks = ["GM12878"],
off_tasks = ["K562"],
model = grelu_model,
compare_func='subtract', # Maximize the difference in probabilities
)
from grelu.design import evolve
output = evolve(
test_input, # The initial sequences
grelu_model,
prediction_transform=gm12878_specificity, # Objective to optimize
max_iter=10, # Number of iterations for directed evolution
num_workers=16,
devices=0,
return_seqs="all", # Return the best sequences in each iteration
batch_size=256,
)
Iteration 0
Best value at iteration 0: 0.015
Iteration 1
Best value at iteration 1: 0.031
Iteration 2
Best value at iteration 2: 0.070
Iteration 3
Best value at iteration 3: 0.113
Iteration 4
Best value at iteration 4: 0.174
Iteration 5
Best value at iteration 5: 0.233
Iteration 6
Best value at iteration 6: 0.282
Iteration 7
Best value at iteration 7: 0.325
Iteration 8
Best value at iteration 8: 0.374
Iteration 9
Best value at iteration 9: 0.419
Iteration 10
Best value at iteration 10: 0.492